Project Summary
| Name | Value |
|---|
Data Download
Download 4901 protein structures predicted for Giardia duodenalis, protein-ligand complexes, and structural homology information
Download
Term Definitions
| Term | Definition |
|---|---|
| AA_%ID | Percentage amino acid identity between query and reference |
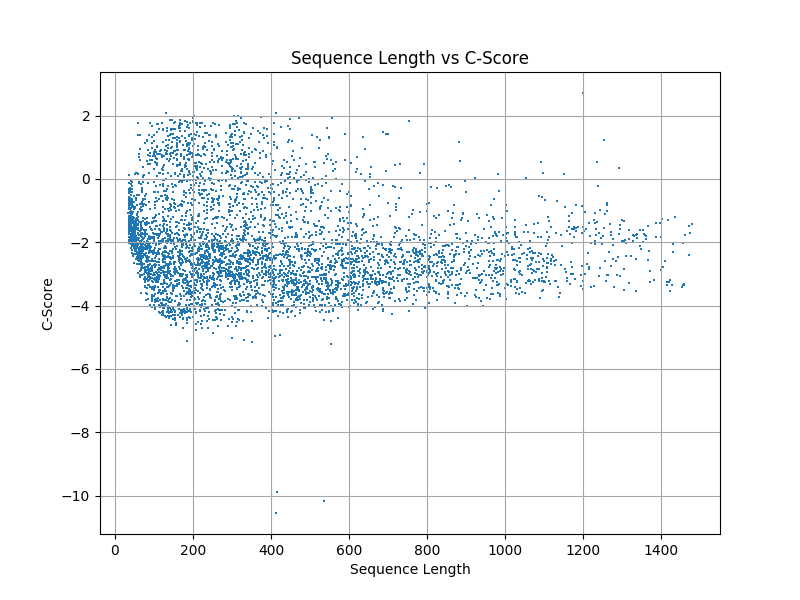
| C_score | Convergence of simulated structures |
| C_sd | Error in C_score |
| Code_Chain | Closest structural homologue in PDB (reference) database |
| Confidence_category | Confidence in model (query) structure based on agreement between actual and predicted PFAM match status.
|
| Cov | Three dimensional coverage of query structure relative to reference |
| Exact_match_prediction | Likelihood of exactMatch between query and reference, output from random forest classifier |
| Length_ratio | Ratio of query peptide AA length to reference peptide AA length |
| Molecule | Description of PDB reference molecule |
| RMSD | Root mean squared deviation in alpha carbon atoms between query and reference |
| RMSD_model | Expected RMSD |
| RMSD_model_sd | Error in RMSD_model |
| Species | Species encoding reference structure |
| SS_sd | Standard deviation of secondary structure proportions (lower indicates greater complexity) |
| TM | Template modeling score - describes overall goodness of fit between query and reference structures |
| TM_model | Expected TM score |
| TM_model_sd | Error in TM_model |